Progress
The New Way to protect. prevent.
We are working to change the world with the first universal influenza vaccine.
Progress to Date
-
02/10
Dr. Arthur Young builds mutant plasmid libraries for all eight flu segments, with every possible single nucleotide position mutated >100x over.
-

09/10
Plasmid libraries are used to produce virus mutant libraries, which are selected by two rounds of infection in cell culture. Sequencing of complex mutant sample after selection by high-throughput, next-generation sequencing (SOLiD).
-
01/11
First-of-kind NGS sequencing error-correction method successfully implemented, and sequencing data analyzed.
-
01/13
Nicholas Wu, graduate student protege of Dr. Young, repeats selection and sequencing utilizing improved molecule (barcode) tagging and using Illumina HiSeq2000 (superior to SOLiD). Obtains deeper coverage from sequencing. After data analysis, vaccine epitope list expanded to 45 peptides.
-
03/13
More than 50 individual mutants validated by reconstruction of mutant and testing in cell culture. Validation rate ~80% overall.
-

10/16
Derived list of 15 epitopes taking the merger of 174 experimentally validated human Influenza A virus epitopes (www.fludb.org) and the most highly invariant epitopes based on our next-gen sequencing data.
-

12/16
Provisional patent filed for the 15 epitopes.
-

05/17
Proof of concept successful: a 4-epitope, adenoviral-vectored vaccine taking the most highly invariant BALB/C mouse flu epitopes achieves 100% protection from lethal flu challenge in mice.
-
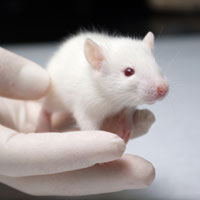
04/18
Mouse (4-epitope) vaccine protects 100% against two strains of lethal H3N2 challenge: X31 and HK68.
-
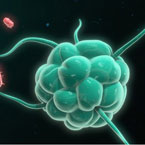
07/18
Intracellular cytokine staining from mice immunized with 4-epitope vaccine shows strong immunogenicity against 2 epitopes and weaker against 2 epitopes.
-

02/19
Detailed experiment establishing preexisting immunity to adenovirus vector shows that, under the right conditions, our adenovirus-vectored flu vaccine is a viable modality in humans with preexisting immunity to naturally circulating adenovirus.
-

05/19
Human (15-epitope) vaccine protects 100% against H1N1 flu in mice.
-

06/19
Mouse vaccine protects 80% against highly pathogenic avian (H5N1) flu in mice, compared to 35% for the standard-of-care (quadrivalent vaccine).
-

11/19
Human vaccine is immunogenic against two HLA-B27 flu peptides in human peripheral blood mononuclear cells in vitro, and two HLA-B27 flu peptides in human dendritic cells in vitro, the best predictor to date of immunogenicity in humans.
-

4/21
Patent for 11 sequence stretches comprised by 48 peptides granted.
-
1/22
National Science Foundation SBIR awarded for a prophylactic antibody project.
-

2/22
Department of Defense Congressionally Directed Medical Research Project grant awarded for workup of Hepatitus B virus genome toward an improved CRISPR/Cas therapy.
-

8/22
Large experiment in ferrets successfully shows statistically significant correlation between T cell activation status and protection from flu challenge.
-
5/23
Second patent broadening claimset for flu peptides granted.
